Protein Design for ML Researchers
Note: This post is written for ML researchers entering the protein design field. It covers the structural biology you need to read RFDiffusion and ProteinMPNN papers, understand what pLDDT and ipTM measure, and follow a design project from target definition through experimental validation. No biology background is assumed.
Introduction
Small-molecule drugs dominate pharmacology, but they have hard limits. A typical small molecule has a molecular weight under 500 Da and contacts its target over ~300 Å\(^2\) (1 Å = 0.1 nm). Proteins are 100–1000x larger, with binding interfaces spanning 1,000–2,000 Å\(^2\). This surface area difference translates to selectivity: proteins can distinguish targets that small molecules cannot, because they encode more geometric and chemical information at the interface.
Many disease-relevant targets — protein–protein interactions, flat surfaces, disordered regions — are “undruggable” by small molecules because there is no deep binding pocket to exploit. Designed proteins can bind these surfaces directly. The prefusion-stabilized COVID-19 spike (Hsieh et al., 2020) was used in multiple approved vaccines, and de novo designed miniprotein inhibitors achieved picomolar (extremely tight) binding to the spike in lab assays (Cao et al., 2020).
For ML researchers, the appeal is structural. Protein design is a well-defined generative modeling problem: the input is a functional specification (target structure, binding constraints), the output is a sequence of discrete tokens (amino acids) that must satisfy continuous geometric constraints (3D folding). The training data is the Protein Data Bank — ~200,000 experimentally solved structures — supplemented by billions of sequences from genomic databases. Structure prediction (AlphaFold) provides a fast oracle for evaluating designs. And the experimental feedback loop is tight: a design campaign from computation to lab results takes weeks, not years.
The field has hit an inflection point. Before 2020, computational protein design relied on Rosetta’s physics-based energy function and Monte Carlo sampling — slow, expensive, and with low success rates. Between 2021 and 2023, AlphaFold2, ProteinMPNN, and RFDiffusion replaced the core steps of the pipeline with learned models, increasing success rates from ~1% to ~10–30% and reducing computation from CPU-weeks to GPU-hours. This created an opportunity: the tools work, but they are far from optimal, and the design space is enormous. Protein design is now a field where ML contributions have immediate, measurable experimental impact.
This post covers the biology and computational infrastructure you need to participate. It follows the sequence → structure → function triangle: what proteins are, what holds them together, what they do, and how we design new ones.
Roadmap
| Section | What It Covers |
|---|---|
| What a Protein Is | Amino acids, structure hierarchy (primary through quaternary), folds, domains, MSAs and coevolution |
| What Holds the Shape Together | The hydrophobic effect, hydrogen bonds, salt bridges, disulfide bonds, stability, and aggregation |
| What Proteins Do | Binding affinity and specificity, interfaces, hotspots, antibodies, enzymes |
| How Protein Designers Think | Energy landscapes, packing, H-bond networks, shape complementarity — bridging biology and ML intuition |
| The Design Problem | Three ML formulations (forward folding, inverse folding, de novo generation), design targets and constraints |
| The Computational Toolkit | Rosetta, AlphaFold, ESMFold, ProteinMPNN, RFDiffusion, Boltz, BoltzGen, and key metrics |
| The Design Workflow | The five-step pipeline from backbone generation to experimental validation |
What a Protein Is
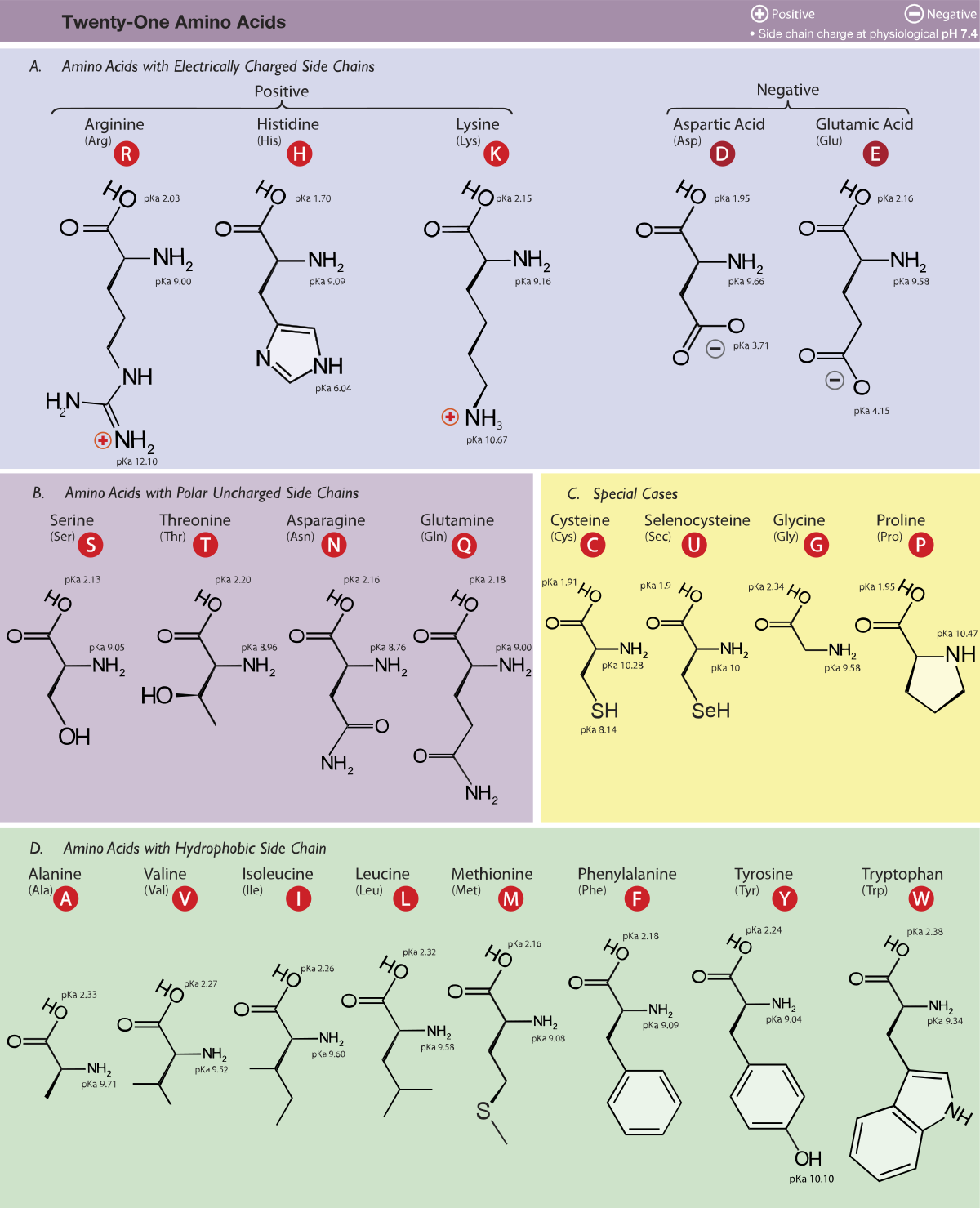
A protein is a chain of amino acids — small molecules linked end-to-end like beads on a string. There are 20 types, each identified by a one-letter code (A for alanine, M for methionine, etc.), so a protein sequence reads like MKVLWAGG... — a string over a 20-letter alphabet.
Every amino acid shares the same backbone atoms — a repeating N-C\(_\alpha\)-C unit — but differs in its side chain, the group that branches off at each C\(_\alpha\). Side chains vary in size, charge, and hydrophobicity.1 This chemical diversity gives proteins their functional range.
Structure hierarchy
Proteins organize at four levels:
- Primary structure — the amino acid sequence itself.
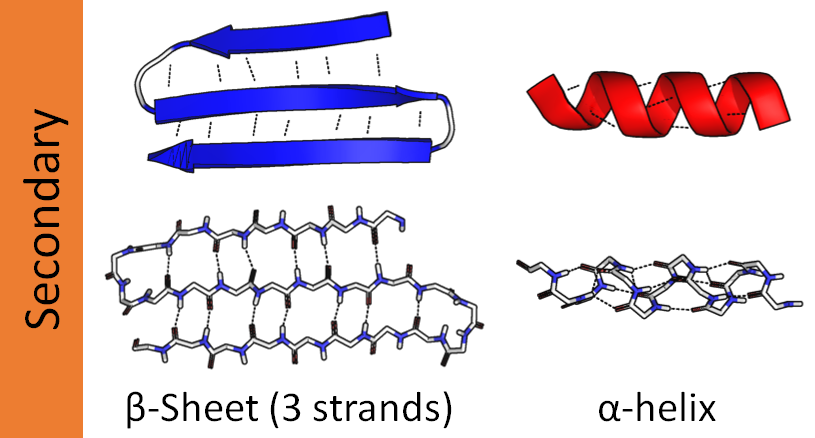
- Secondary structure — local repeating patterns. Alpha helices are coiled springs stabilized by hydrogen bonds between every 4th residue (a “residue” is one amino acid in the chain). Beta sheets are flat arrangements of adjacent strands connected by hydrogen bonds. Loops are the flexible connectors between them.
- Tertiary structure — the full 3D shape of a single chain, with helices, sheets, and loops packed together.
- Quaternary structure — the assembly of multiple chains into a complex.
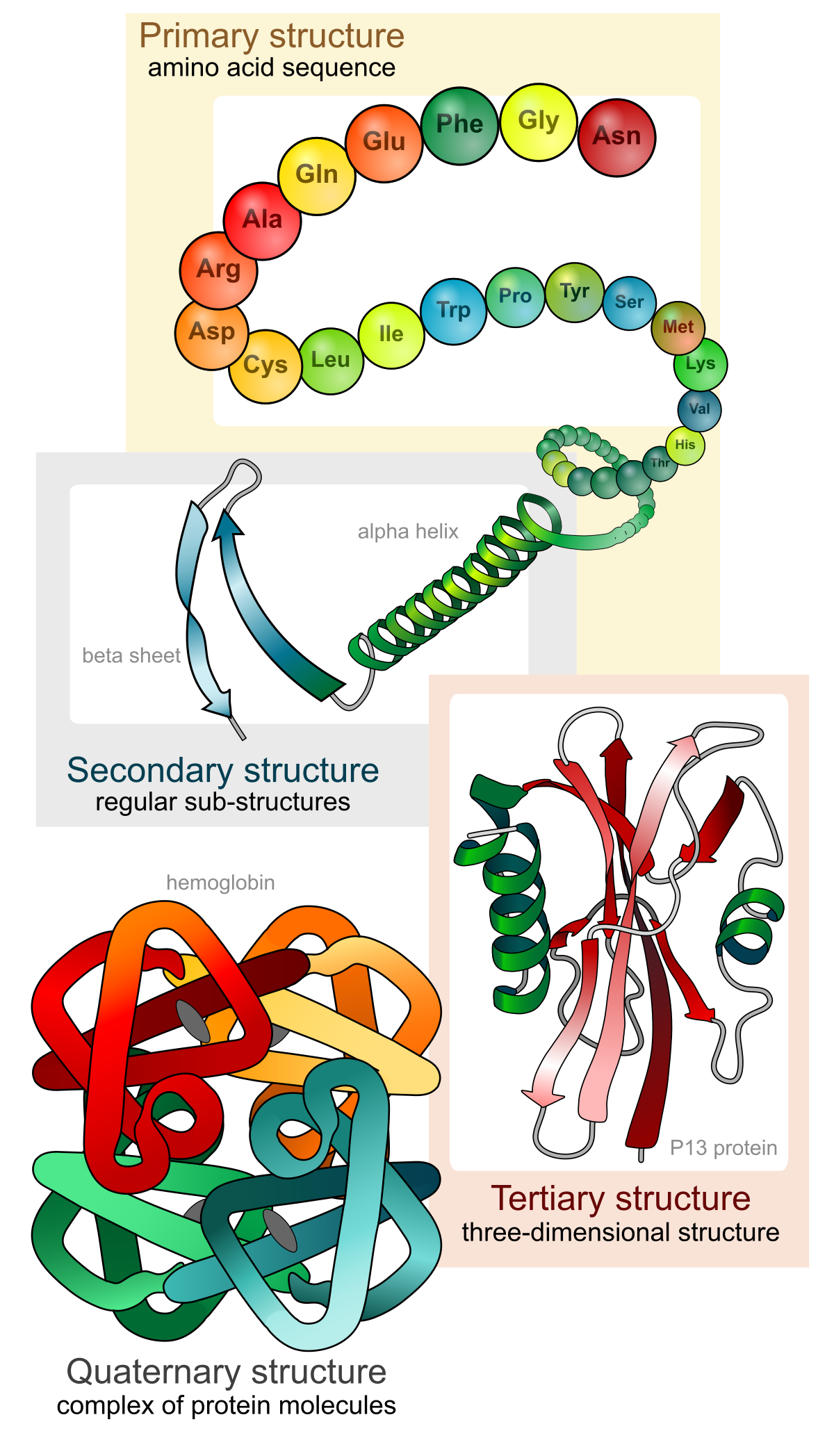
A fold (or topology) is the overall arrangement of secondary structure elements. Two proteins with completely different sequences can share the same fold — different bricks, same floor plan. A domain is a compact, independently folding unit within a larger protein; many proteins consist of multiple domains linked together.
Evolution and MSAs
Related proteins across species — homologs — share a common ancestor. Lining up homologous sequences produces a multiple sequence alignment (MSA), which reveals conservation: positions that stay constant across millions of years of evolution are structurally or functionally critical.

MSAs also reveal coevolution — pairs of positions that mutate together, implying physical contact. This was the key insight behind early contact prediction methods and a core input to AlphaFold2. For ML researchers, MSAs are the protein equivalent of a large unlabeled dataset: they encode structural constraints without explicit 3D labels.
The organizing principle of structural biology is the sequence → structure → function triangle: sequence determines the 3D fold, and the fold determines what the protein does.
What Holds the Shape Together
A protein folds because it is thermodynamically favorable to do so. The dominant driving force is the hydrophobic effect: nonpolar side chains are energetically penalized when exposed to water, so the chain collapses to bury them in a tightly packed interior — the hydrophobic core. Disrupting this core usually destroys the protein.
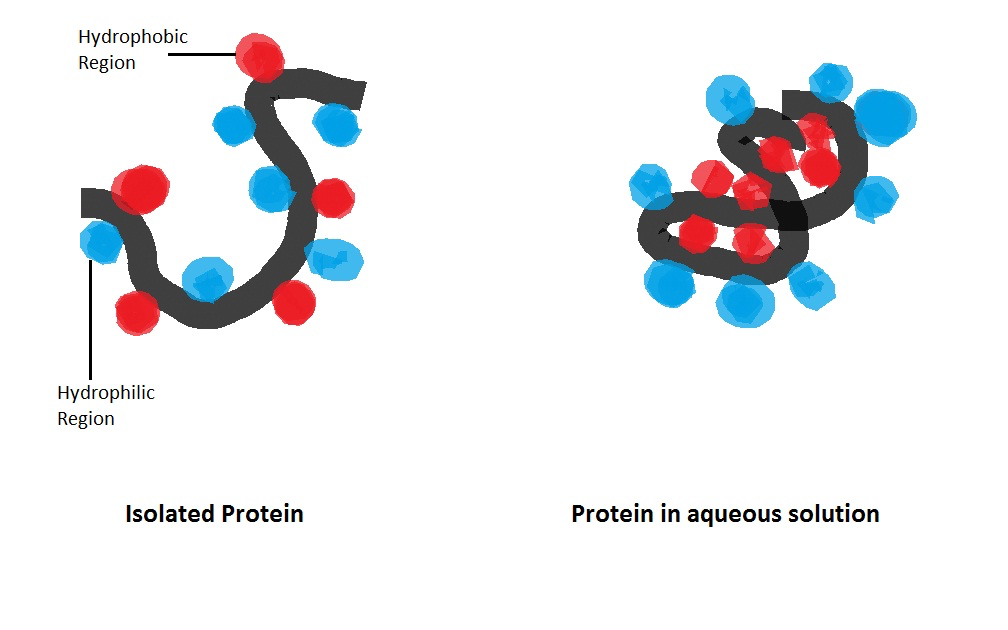
On top of the hydrophobic effect, several other forces contribute:
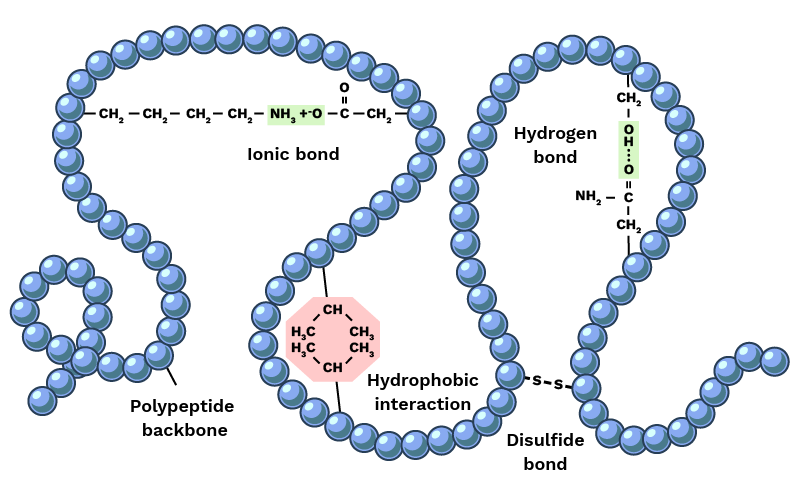
- Hydrogen bonds — weak electrostatic attractions between donor and acceptor atoms. Individually modest, but hundreds of them collectively define secondary structure. Every backbone N-H and C=O must either form an H-bond or be exposed to water; an unsatisfied H-bond donor buried in the core is energetically catastrophic.
- Salt bridges — attractions between positively charged residues (Lys, Arg) and negatively charged ones (Asp, Glu). Contribute to stability on the protein surface.
- Disulfide bonds — covalent bonds between two cysteine residues. Molecular staples that physically lock distant parts of the chain together. Common in antibodies and secreted proteins.
- Van der Waals interactions — weak attractions between atoms at close range. Negligible individually but significant when combined, as thousands of atoms pack tightly in the core.
Stability and failure modes
Stability measures how hard it is to unfold a protein. The melting temperature (T\(_m\)) is the temperature at which half the protein population is unfolded — higher T\(_m\) means a more robust protein. Designed proteins typically need T\(_m\) > 60°C to be useful.2
The most common failure mode in protein design is aggregation: proteins stick to each other and form useless clumps, like egg whites cooking. This usually happens because hydrophobic patches that should be buried are instead exposed on the surface. Solubility — whether the protein stays dissolved in water — is closely related. An insoluble protein is useless regardless of how good it looks in simulation.
What Proteins Do — Binding and Function
Most proteins function by binding to other molecules — other proteins, small molecules, DNA, or metal ions. The strength of binding is quantified by the dissociation constant K\(_d\):3 lower K\(_d\) means tighter binding (the two molecules are harder to pull apart).
| K\(_d\) range | Binding strength | Typical context |
|---|---|---|
| < 1 nM | Very tight | Therapeutic antibodies |
| 1–100 nM | Tight | Designed binders, drugs |
| 100 nM – 1 μM | Moderate | Signaling interactions |
| > 1 μM | Weak | Transient contacts |
Specificity is equally important: a binder that grabs everything is useless. The physical contact surface between two binding partners is the binding interface, typically spanning 1,000–2,000 Å\(^2\). Not all interface residues contribute equally — a handful of hotspot residues provide most of the binding energy. Identifying hotspots on the target surface is the first step of binder design.
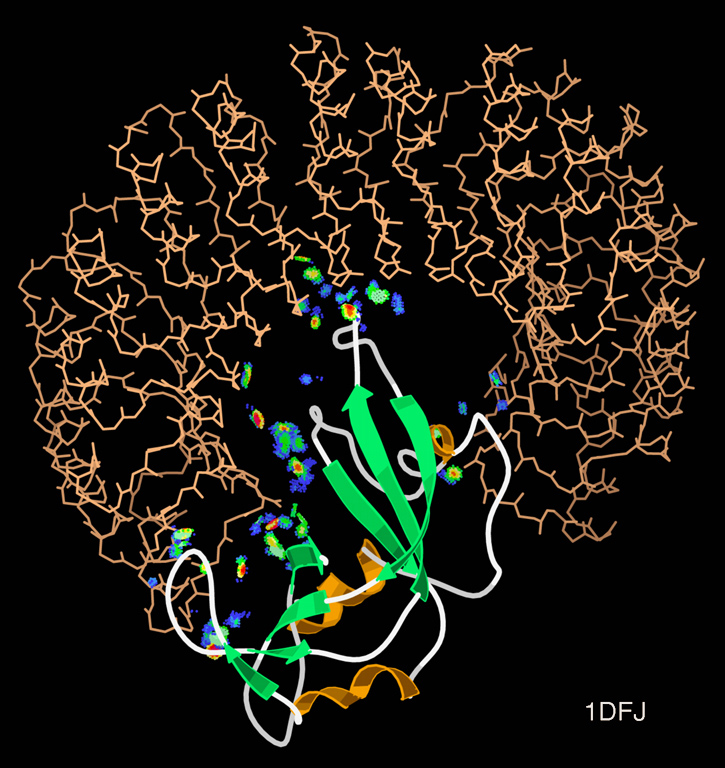
For antibodies specifically, the target surface is called the epitope and the matching surface on the antibody is the paratope. Different antibodies can target different epitopes on the same target protein.
Beyond binding, enzymes are proteins that catalyze chemical reactions. Their active sites — small pockets with precisely positioned catalytic residues — accelerate reactions by factors of 10\(^6\)–10\(^{12}\). Enzyme design is harder than binder design because it requires exact 3D geometry, not just a good surface fit.
Proteins can also undergo conformational changes — shifts in 3D structure triggered by binding. This is how signals propagate through biological systems: binding at one site rearranges the protein to expose or hide a distant functional site.
How Protein Designers Think
ML researchers sometimes imagine that protein designers understand proteins through deep biological intuition — years of staring at crystal structures, building up an indescribable feel for what works. The reality is more specific than that. Protein designers think in rules. When they look at a structure, they ask concrete questions: Are the hydrophobic residues buried? Are the hydrogen bond donors satisfied? Is the backbone in a favorable region of Ramachandran space?4 Much of their expertise is not mystical pattern recognition — it is a mental checklist of physical constraints.
This is good news for ML researchers, because many of these heuristics translate naturally into energy function terms, loss functions, and model inductive biases.
Energy landscapes
The folded state of a protein sits at the bottom of a free energy landscape — a surface over the space of all possible conformations. Design means finding sequences whose energy minimum matches a target structure — an optimization problem.
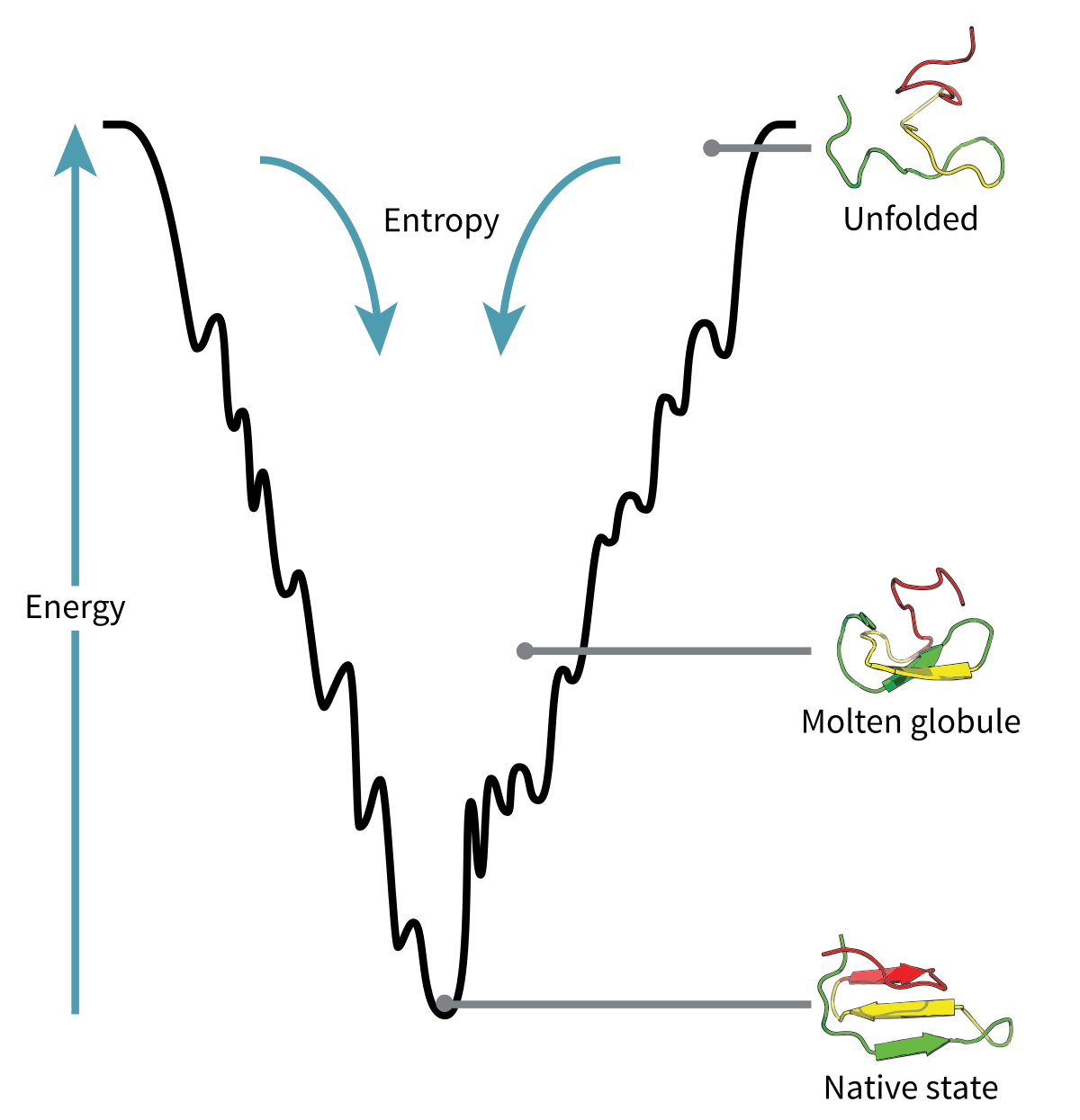
Packing geometry
When a designer says “the hydrophobic core is well-packed,” they mean atoms fill space tightly with no voids. This is quantified by packing density metrics and reflected in the van der Waals energy term in Rosetta. Empty space in the core means unfavorable energetics.
Hydrogen bond networks
“Satisfying all hydrogen bonds” is not vague — it means every backbone and side-chain donor/acceptor that is buried must have a partner. An unsatisfied H-bond donor buried in the core costs roughly 5 kcal/mol, enough to destabilize the entire protein. Designers check these systematically.
Shape complementarity
Binding interfaces are scored by geometric fit using the Sc score, which measures how well the two surfaces fit together (like interlocking fingers). Sc = 1.0 is a perfect fit; most natural protein–protein interfaces score 0.6–0.7. This is a geometric computation, not a subjective judgment.
Systematic enumeration
Rosetta’s design protocol is Monte Carlo sampling over sequence space with a physics-based energy function — an optimization algorithm. At each position, Rosetta tries different amino acid identities and side-chain rotamers,5 accepts or rejects changes based on the energy function, and iterates. The transition from Rosetta to ML-based design was not a paradigm shift in thinking — it was swapping the optimizer from physics-based MCMC to learned models.
Biologist reasoning is ML reasoning
Three examples of “structural biology intuition” translated to ML terms:
| What a biologist says | What they mean computationally |
|---|---|
| “This helix should be amphipathic (hydrophobic on one side, hydrophilic on the other)” | The hydrophobic moment vector should point inward — a periodic constraint on sequence hydrophobicity with period 3.6 (the helix repeat) |
| “The core isn’t packed well” | Atoms have too much empty space — packing density below threshold, Rosetta vdW energy too high |
| “That loop will be floppy” | High B-factor in crystal structures / low pLDDT in AlphaFold — the model is uncertain about this region’s conformation |
The domain knowledge that ML researchers admire in structural biologists is a set of quantitative constraints and heuristics that map to loss function terms and architectural priors. When a biologist says something about a structure, there is usually a computable quantity behind it.
The Design Problem — Formulations and Types
Protein design decomposes into three ML problem formulations:
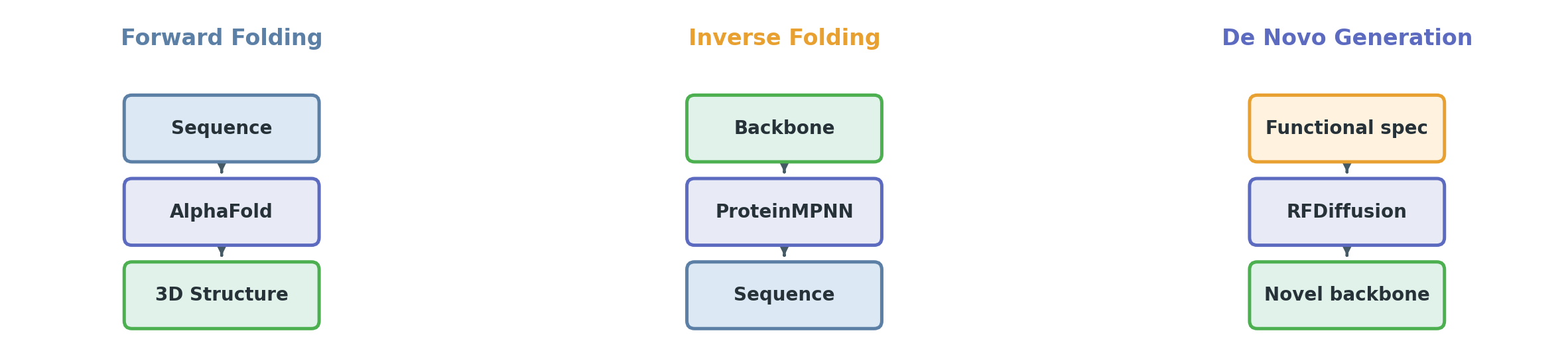
Forward folding (structure prediction). Input: amino acid sequence. Output: 3D atomic coordinates. Model: AlphaFold2. This is the “check your work” step — given a designed sequence, does the predicted fold match what you intended?
Inverse folding (sequence design). Input: 3D backbone coordinates. Output: amino acid sequence that folds into that backbone. Model: ProteinMPNN. This is the core design step — “I drew the blueprint, now find the bricks.”
De novo backbone generation. Input: functional specification (target protein, hotspot residues, symmetry constraints). Output: new backbone structure. Model: RFDiffusion. This is the generative step — “create a new shape that binds to this target.”
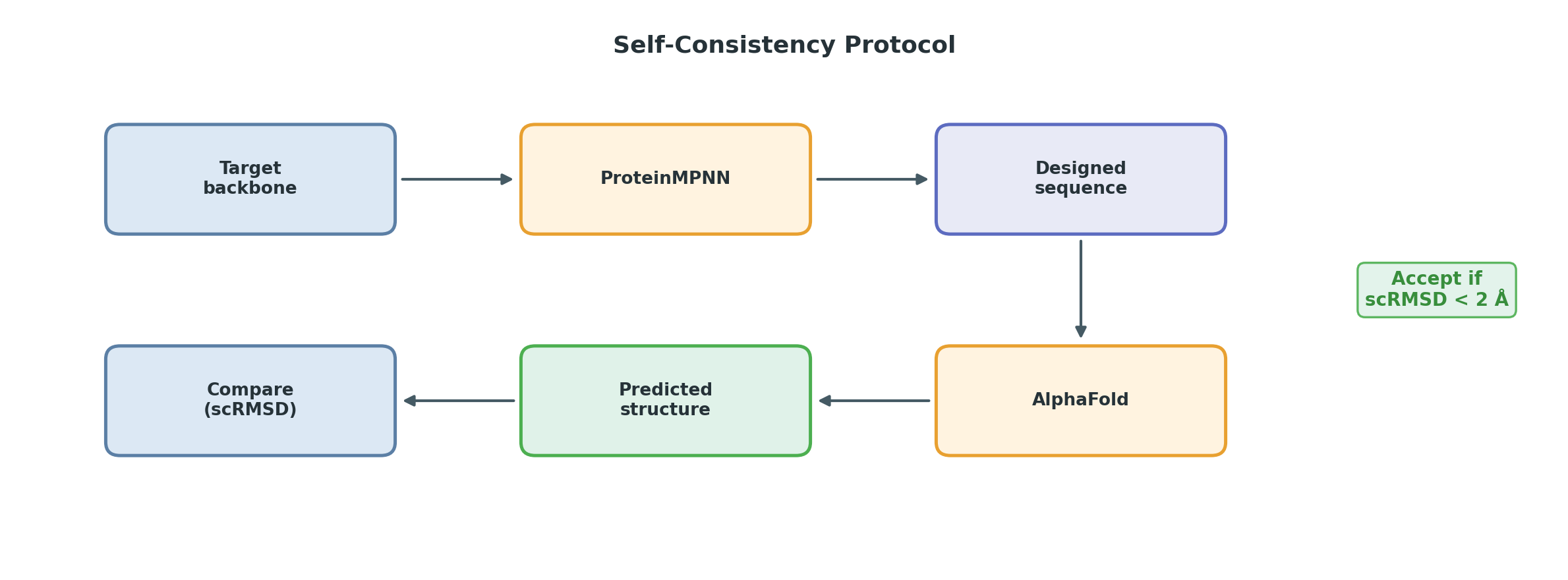
The standard validation loop ties these together: generate a backbone (RFDiffusion) → design a sequence for it (ProteinMPNN) → predict the structure of that sequence (AlphaFold) → compare the prediction to the intended backbone. If they match, the design is self-consistent.
What people design
Design targets:
- Miniproteins (40–80 residues) — small, stable, easy to produce. The “Hello World” of protein design.
- De novo binders — proteins that grab a specific target surface. The most active area in computational design.
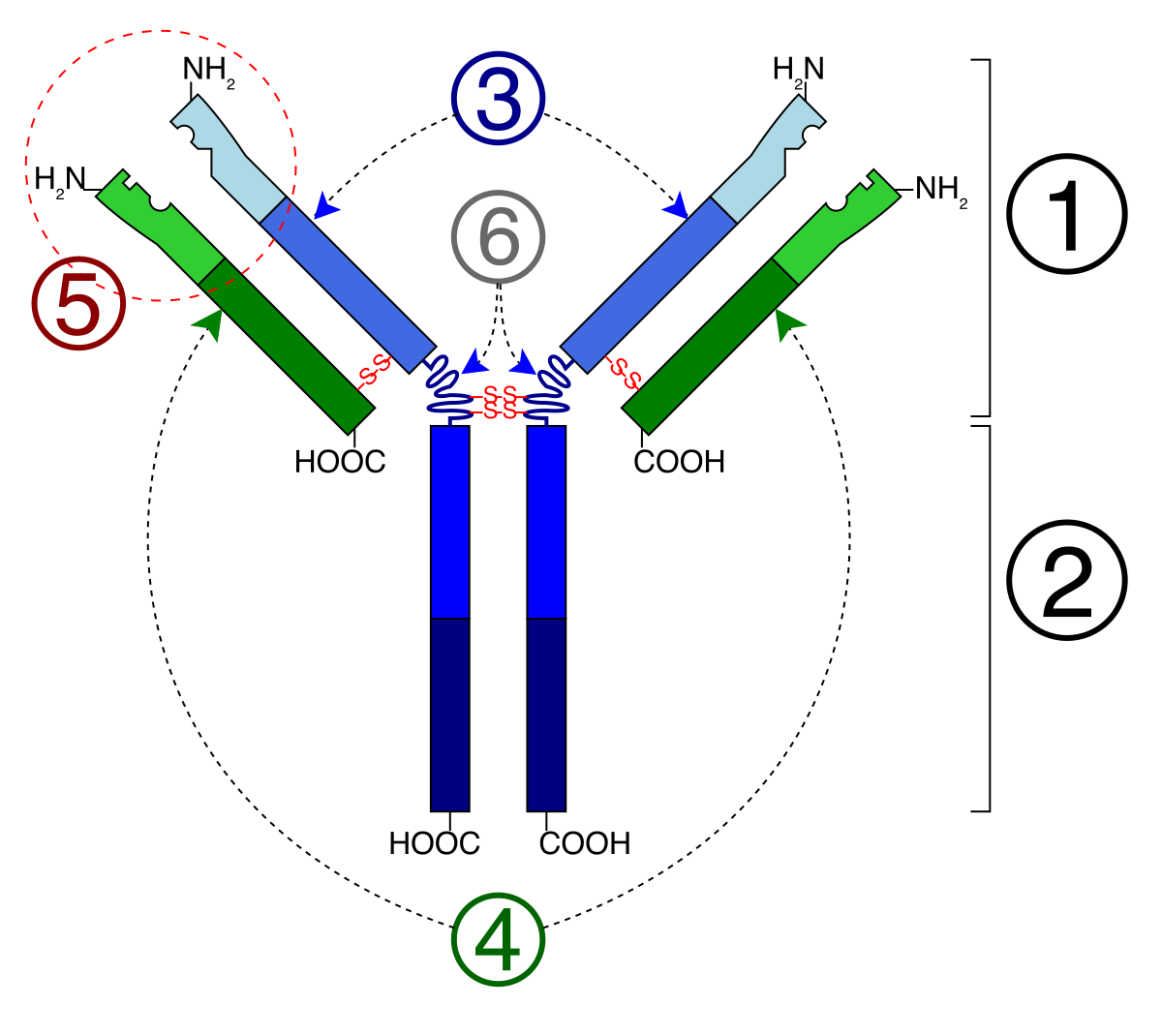
- Antibodies — Y-shaped immune proteins. ~100 approved as drugs. Design focuses on the six CDR loops (especially CDR-H3), which contact the antigen. The stable framework regions hold the CDRs in position.
- Nanobodies (VHH) — single-domain antibodies from camelids (camels, llamas). Smaller, simpler to engineer computationally.
- Peptides (< 40 residues) — short, often flexible chains. Many drugs are peptides (e.g., insulin).
- Enzymes — proteins that catalyze chemical reactions. Harder than binder design: requires precise 3D geometry at the active site.
- Vaccine immunogens — engineered proteins that train the immune system. The COVID-19 spike protein vaccines are a high-profile example.
Design constraints
Real designs are constrained:
- Hotspot residues — “the designed binder must contact these specific residues on the target.”
- Motif scaffolding — “build a stable protein around this functional fragment, holding it in the correct 3D position.”
- Symmetric design — “generate identical subunits that assemble into a ring, cage, or icosahedron (a 20-faced sphere-like shell).”
- Contig notation — RFDiffusion’s input format. Example:
A1-100/0 30-50/B1-80means “keep residues 1–100 of chain A, design 30–50 new residues, keep residues 1–80 of chain B.”
The Computational Toolkit
Seven tools define the current protein design stack. For each: what goes in, what comes out, and when to use it.
Rosetta
The classic physics-based suite, developed over 20+ years. Rosetta evaluates designs using an energy function that sums van der Waals packing, electrostatics, hydrogen bonds, solvation, and backbone geometry terms. Output: Rosetta Energy Units (REU) — lower is better. Still widely used for scoring and refinement, even as ML tools handle generation.
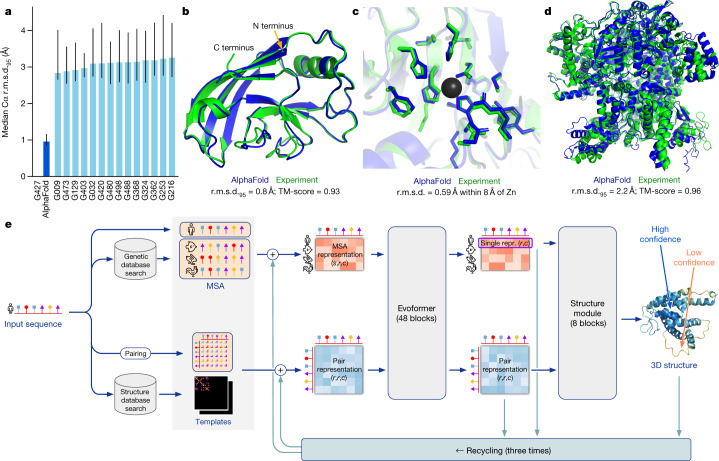
AlphaFold2 / AlphaFold3
Input: sequence (+ MSA for AF2). Output: predicted 3D structure with per-residue and per-pair confidence scores.
- pLDDT — per-residue confidence (0–100). High pLDDT means the model is certain about local structure.
- pTM — overall fold confidence.
- ipTM — interface confidence for protein complexes. The key metric for binder design.
- PAE — predicted aligned error matrix. Shows expected positional error between all residue pairs. For binder design, check the inter-chain PAE block: low values mean the model is confident about the binding mode.
AF2 (with AF2-Multimer) handles single chains and multi-chain protein complexes. AF3 extends to protein–nucleic acid and protein–small molecule complexes.
ESMFold / ESM3
Meta’s protein language models. ESMFold predicts structure from a single sequence — no MSA required, making it much faster than AlphaFold. ESM3 is more general, jointly modeling sequences, structures, and functional annotations. Used for rapid self-consistency screening at scale: when you have 80,000 candidate sequences, you filter with ESMFold first and send only the top candidates to AlphaFold.
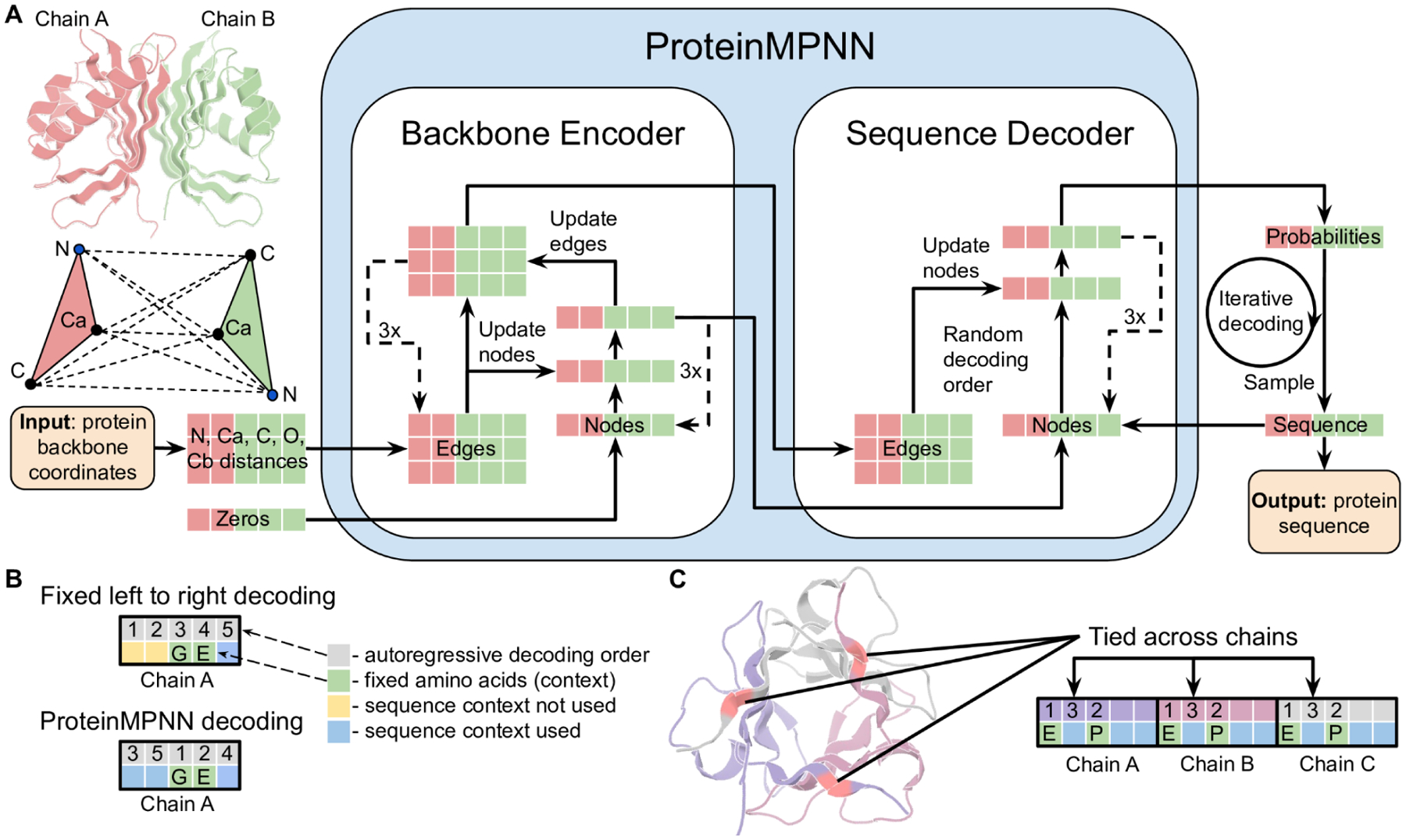
ProteinMPNN
Input: 3D backbone coordinates (as a graph of residue positions). Output: amino acid probability distribution at each position. A message-passing neural network that designs sequences for given backbones. Fast, accurate, and the standard inverse folding tool. Typically generates 8–16 sequences per backbone.
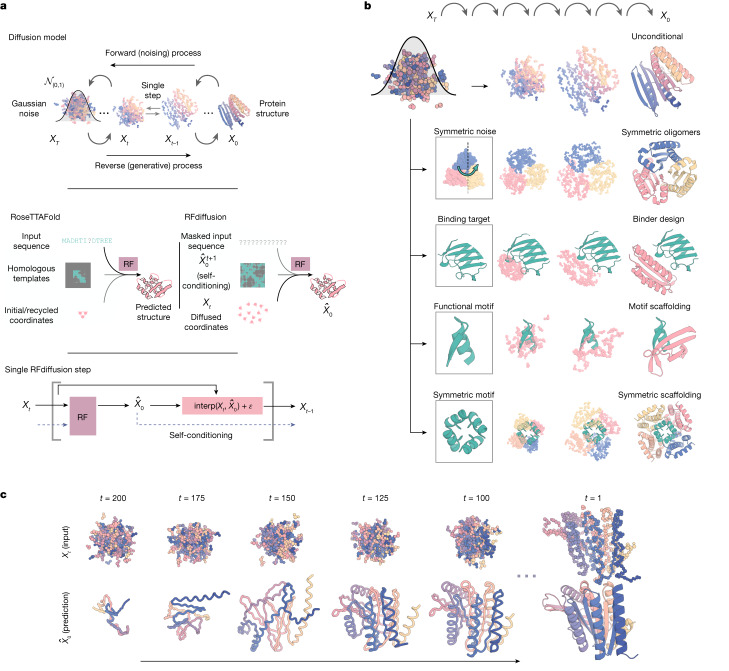
RFDiffusion
Input: target protein structure + conditioning constraints (hotspot residues, motifs, symmetry). Output: new backbone coordinates. A denoising diffusion model over backbone coordinates — analogous to image diffusion models but operating in SE(3) coordinate space. The most powerful current tool for de novo backbone generation.
Boltz / Boltz2
Alternative structure predictors that provide a second opinion on AlphaFold. Useful when you want independent confirmation that a designed sequence folds correctly. Different architecture, different training data, so agreement between AlphaFold and Boltz is a stronger signal than either alone.
BoltzGen
The standard pipeline above — RFDiffusion for backbones, ProteinMPNN for sequences, AlphaFold for validation — treats each step as a separate model. BoltzGen (Stark et al., 2025) integrates these into a unified pipeline that co-designs sequence and structure.
Input: target structure + design specification (binding site, covalent constraints, sequence length range). Output: designed binder with both 3D coordinates and amino acid sequence. BoltzGen’s pipeline has several stages beyond the core generative model:
- Structure generation — an all-atom diffusion model generates candidate binder structures conditioned on the target.
- Solubility-aware inverse folding — a separate inverse folding model, trained on soluble proteins, re-sequences the generated structures to improve solubility and expression. This addresses a common failure mode: designs that look good computationally but don’t express in bacteria.
- Co-folding validation — Boltz predicts the complex structure from the designed sequence to check self-consistency.
- Quality filtering — physics-based metrics are computed for each design: number of hydrogen bonds and salt bridges at the interface, change in solvent-accessible surface area upon binding, and surface hydrophobicity. Each design is ranked on every metric, and the worst rank across all metrics becomes the overall quality score.
- Quality-diversity selection — a greedy algorithm picks the final candidates by iteratively choosing designs that score high and are structurally distinct from those already selected. This prevents the final set from collapsing to minor variants of one solution.
The pipeline supports design types that the modular RFDiffusion/ProteinMPNN stack handles poorly: cyclic peptides, disulfide-bonded peptides, and nanobodies.
Experimental validation across 8 wet-lab campaigns showed nanomolar binders for 66% of novel protein targets (less than 30% sequence identity with any known bound structure in the PDB).
What does “good” look like?
A reference table for interpreting computational metrics:
| Metric | Good threshold | What it means |
|---|---|---|
| pLDDT | > 80 | High confidence in local structure |
| ipTM | > 0.8 | Confident interface prediction |
| scRMSD | < 2 Å | Self-consistent design (sequence encodes the intended fold) |
| K\(_d\) | < 100 nM | Tight binding (experimental measurement) |
| Rosetta energy | Negative, comparable to natural proteins | Physically reasonable packing and interactions |
The Design Workflow
The tools above assemble into a five-step pipeline. The key insight is the numbers at each stage — the funnel from millions of computational candidates to a handful of experimental hits shapes every decision in a design project.
Step 1: Define the problem
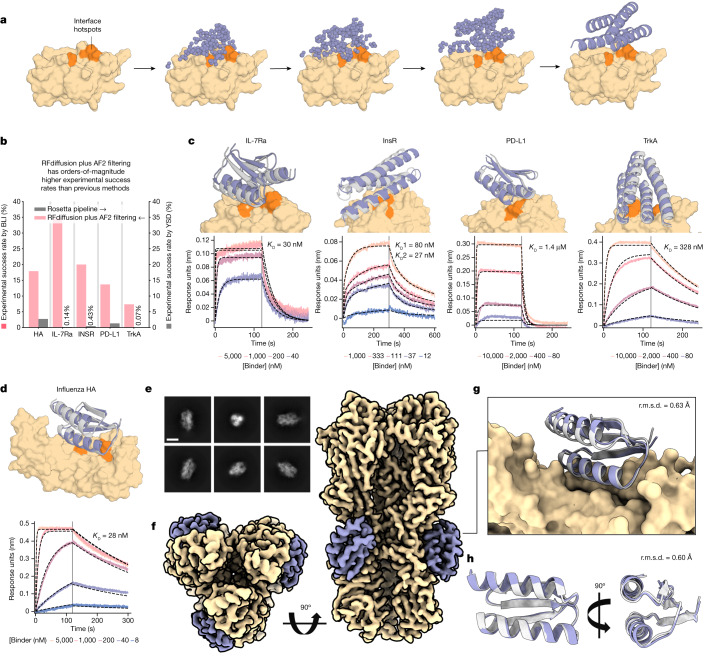
What should the protein do? Bind a specific target surface? Catalyze a reaction? Form a symmetric cage? This determines which tools to use, which constraints to set, and how to evaluate success. For binder design, you identify the target protein, choose an epitope (the surface patch to target), and specify hotspot residues.
Step 2: Generate backbones
RFDiffusion generates ~10,000 backbone structures conditioned on the target and constraints. Each backbone is a candidate protein shape — no sequence yet, just the 3D arrangement of backbone atoms.
Step 3: Design sequences
ProteinMPNN designs 8–16 amino acid sequences for each backbone, yielding ~80,000–160,000 total candidates. Each is designed to fold into the target backbone.
Step 4: Computational filtering
The self-consistency check eliminates most candidates. For each designed sequence:
- Predict its structure with AlphaFold or ESMFold.
- Compare the predicted structure to the intended backbone (scRMSD).
- Check confidence metrics (pLDDT, ipTM for binders).
~1% of designs pass all filters — about 800 candidates from 80,000.
Step 5: Experimental validation
The top ~20 candidates are ordered as synthetic genes and tested in the lab:
- Expression — bacteria (usually E. coli) produce the designed protein from synthetic DNA. Many designs fail here: the protein doesn’t express or is insoluble.
- Purification — the protein is isolated from the bacterial cell contents. Aggregated or insoluble proteins can’t be purified.
- CD (circular dichroism) — a quick test for secondary structure. If the CD spectrum shows no helices or sheets, the protein didn’t fold.
- SPR (surface plasmon resonance) — measures binding affinity in real time, providing the K\(_d\) value. The gold standard for quantifying binder quality.
- Display methods (phage display, yeast display) — screen millions of variants simultaneously. Each variant is displayed on the surface of a phage or yeast cell, washed over the target, and only binders stick. Used to improve initial hits.
- Directed evolution — the pre-ML baseline. Randomly mutate, test, keep the best, repeat. Won the 2018 Nobel Prize in Chemistry. Computationally, this is what evolutionary algorithms replicate.
- Cryo-EM / X-ray crystallography — determine the actual 3D atomic structure. The definitive validation: did the protein fold into the intended shape? Slow and expensive, reserved for the most promising candidates.
Typical hit rates: ~10 of 20 ordered designs express as soluble protein, and 3–5 bind the target. These numbers vary by target difficulty, but the order of magnitude is consistent across published studies.
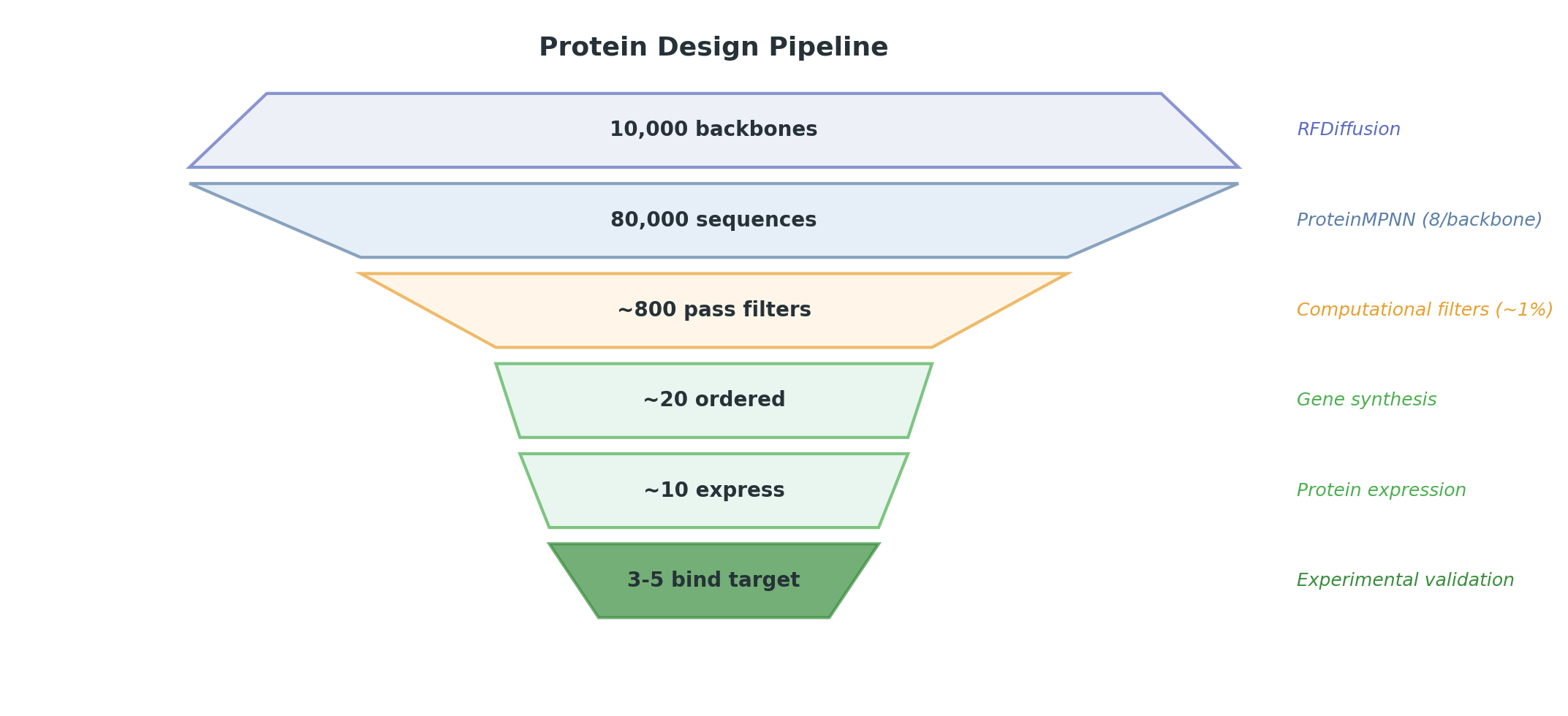
The gap between computational output (millions of candidates) and experimental throughput (tens to hundreds) is the defining constraint of protein design. Every ML improvement that increases the pass rate at Step 4 reduces the cost and time of experimental campaigns. This is where the field’s leverage lies.
References
- Cao, L., et al. (2020). De novo design of picomolar SARS-CoV-2 miniprotein inhibitors. Science, 370(6515), 426–431.
- Hsieh, C.-L., et al. (2020). Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science, 369(6510), 1501–1505.
- Leman, J. K., et al. (2020). Macromolecular modeling and design in Rosetta: recent methods and frameworks. Nature Methods, 17, 665–680.
- Jumper, J., et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596, 583–589.
- Dauparas, J., et al. (2022). Robust deep learning–based protein sequence design using ProteinMPNN. Science, 378(6615), 49–56.
- Lin, Z., et al. (2023). Evolutionary-scale prediction of atomic-level protein structure with a language model. Science, 379(6637), 1123–1130.
- Watson, J. L., et al. (2023). De novo design of protein structure and function with RFdiffusion. Nature, 620, 1089–1100.
- Abramson, J., et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature, 630, 493–500.
- Wohlwend, J., et al. (2024). Boltz-1: Democratizing Biomolecular Interaction Modeling. arXiv:2408.00537.
- Stark, H., et al. (2025). BoltzGen: Toward Universal Binder Design. bioRxiv:2025.11.20.689494.
Figure sources
- Amino acids diagram: Wikimedia Commons, CC BY-SA 3.0.
- Protein structure levels: Wikimedia Commons, public domain.
- Secondary structure (alpha helix and beta sheet): Wikimedia Commons, CC BY-SA 4.0.
- Hydrophobic interaction: Wikimedia Commons, CC BY-SA 3.0.
- Protein interactions: Labster Theory.
- Protein–protein interface (PDB 1DFJ): Wikimedia Commons, CC BY 3.0.
- Antibody structure: Wikimedia Commons, CC BY-SA 3.0.
- Multiple sequence alignment: Wikimedia Commons, CC BY-SA 3.0.
- Folding energy landscape: Wikimedia Commons by Thomas Splettstoesser, CC BY-SA 3.0.
- AlphaFold overview: Figure 1 from Jumper et al. (2021), via PMC8371605.
- ProteinMPNN overview: Figure 1 from Dauparas et al. (2022), via PMC9997061.
- RFDiffusion overview: Figure 1 from Watson et al. (2023), via PMC10468394.
- Binder design examples: Figure 6 from Watson et al. (2023), via PMC10468394.
-
The 20 amino acids split roughly into four groups: nonpolar/hydrophobic (A, V, L, I, M, F, W, P), polar uncharged (S, T, N, Q, Y, C, G), positively charged (K, R, H), and negatively charged (D, E). The nonpolar residues drive folding by burying themselves away from water. ↩
-
T\(_m\) is measured by heating the protein while monitoring secondary structure (e.g., circular dichroism). A well-designed miniprotein might reach T\(_m\) > 90°C. ↩
-
K\(_d\) is the concentration at which half the binding sites are occupied at equilibrium. It has units of molar concentration. K\(_d\) = 1 nM means the binder holds on tightly even at very low concentrations; K\(_d\) = 1 μM is moderate affinity typical of transient interactions. ↩
-
The Ramachandran plot charts the two backbone dihedral angles (\(\phi\), \(\psi\)) for each residue. Some angle combinations cause atomic clashes and are forbidden. A well-designed protein has all residues in the “allowed” regions of this plot. ↩
-
A rotamer is a preferred side-chain conformation. Side chains don’t rotate freely — they snap into a discrete set of low-energy angles, like a dial with set positions. Rosetta’s rotamer library catalogs these preferred conformations for each amino acid type. ↩